Please select your location and preferred language where available.
Real-time extraction of neural voltage from microscopy video
February 7, 2024
Kioxia is developing high-speed microscopy image processing methods leveraging SSDs*. Here we introduce one of the outcomes, which is a method able to extract neural voltage from microscopy video in real time[1].
Inside the brain, neurons exchange electric signals with each other, and studying brain activities by measuring voltage of individual neurons is one of the important means for understanding how the brain works. Traditionally, neural voltage has been measured by placing electrodes or pipettes in physical contact with neurons of experimental animals such as mice, but these devices make it difficult to deal with many neurons at the same time. Moreover, they also damage the brain as foreign objects have to be inserted into the brain. To address these issues, recent years have seen significant technological advances in so-called voltage imaging, where fluorescent materials called voltage indicators are designed to convert neural voltage to light so it can be captured by a microscope[2].
In a nutshell, active neurons appear blinking under the microscope. While there are still impacts on the environment of the brain such as drilling a hole in the skull, voltage imaging enables less invasive measurement of multiple neurons without damaging tissue around the neurons of interest. However, in order to keep track of millisecond-order voltage changes, microscopy video is recorded at from hundreds to thousand frames per second (fps).
This poses a challenge in data processing, as even a ten-second capture can lead to gigabytes of video data. According to the neuroscientists from multiple universities we work with, they wish to conduct animal experiments with different conditions leading to 3-4 hours of measurement in total per day. But since it would take a few days to process resultant 1 terabytes of video data, they have to live with smaller-scale experiments in reality: large-scale experiments would only accumulate data, as data processing would not catch up.
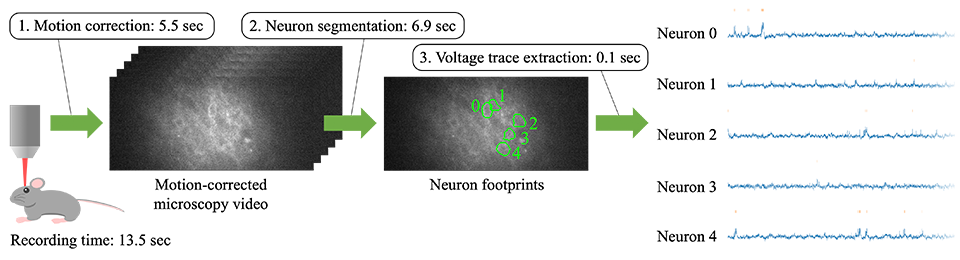
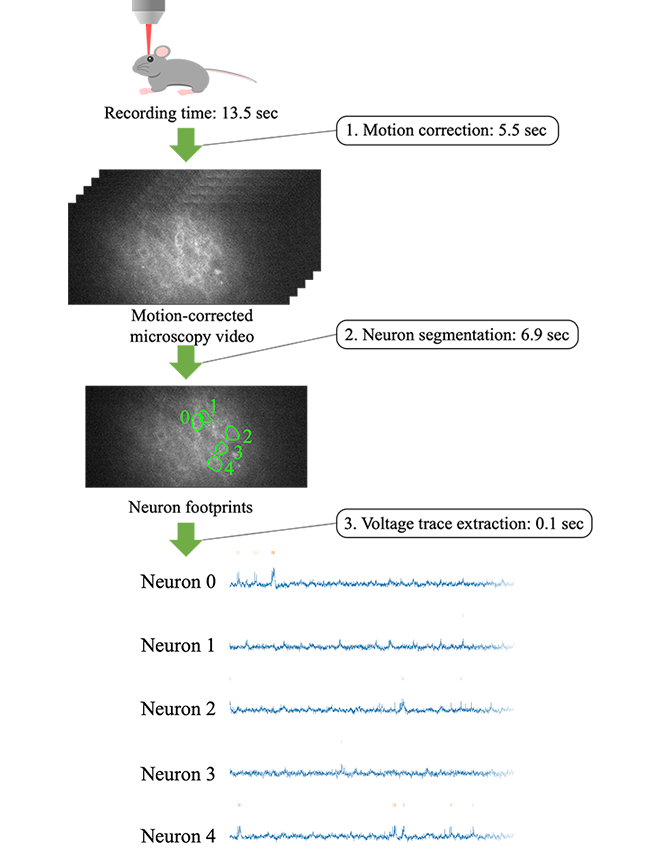
Therefore, Kioxia accelerated the data processing of neural voltage extraction from microscopy video, achieving a processing time that is less than the time it takes to capture the video. This amounts to real-time performance, able to process video as fast as the video capture speed, and thus there will be no longer data that only accumulates without being processed. The data processing pipeline consists of the following three steps (Figure 1).
- Motion correction: Even though the animal is head-fixed to a rig, minute movement manifests itself as non-negligible motion under the microscope. We cancel it via image processing, resulting in video where stationary neurons are blinking.
- Neuron segmentation: We detect neurons from the motion-corrected video and identify the footprint of each neuron in the image.
- Voltage trace extraction: We extract voltage traces from the identified neuron footprints.
The example shown in Figure 1 is a 13.5-second video consisting of 10,000 frames captured at 714 fps. Our pipeline corrects motion in 5.5 seconds, segments neurons (five of them in this example) in 6.9 seconds, and extracts a voltage trace from each neuron in 0.1 seconds. In total, the processing time is 12.5 seconds, which is shorter than the video recording time (or faster than the video recording speed), implying real-time performance.
Step 1 is accelerated through carefully-engineered GPU (graphics processing unit) implementation of well-known existing techniques. They include dividing the image into multiple patches and processing them in parallel on the GPU, as well as table lookup to speed up the summation of pixel values necessary for computing the ZNCC (zero-mean normalized cross-correlation) metric to assess the similarity between frames[3].
Step 2 is designed as shown in Figure 2. First, the motion-corrected input video is temporally split into fixed-length (50 frames) time segments, which are then decimated using summary filters. Each time segment of 50 frames reduces to two summary images, one capturing spatial and the other temporal features, thereby significantly decreasing the number of frames. Subsequently, multiple patches are taken from each summary image pair and fed into the U-Net[4], a convolutional neural network for image segmentation, which outputs a probability representing whether each pixel is a neuron or not. We trained the U-Net using synthetic images simulating the process of microscopy image capture in voltage imaging.
Once trained, the U-Net works fast. By merging patches output by the U-Net, we obtain a probability map of neurons for each time segment. Finally, NMF (nonnegative matrix factorization) is used to decompose the sequence of probability maps into neuron footprints and their temporal profiles.
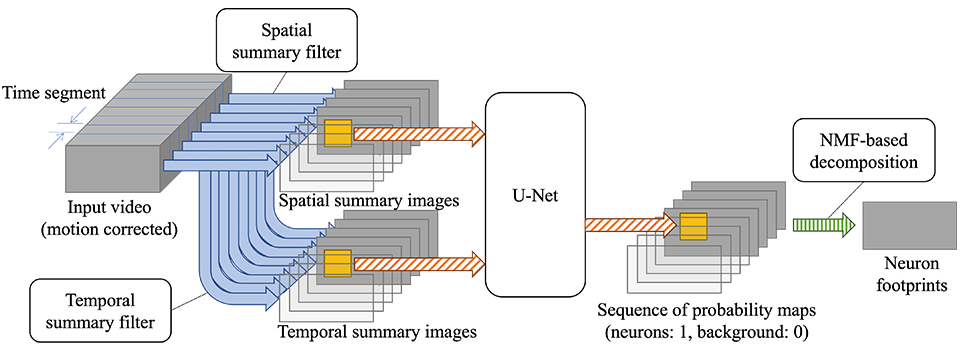
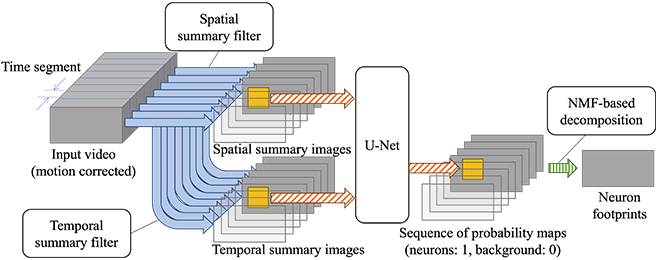
Step 3 simply averages pixel values within each neuron footprint every frame, which is fast enough with straightforward parallelization.
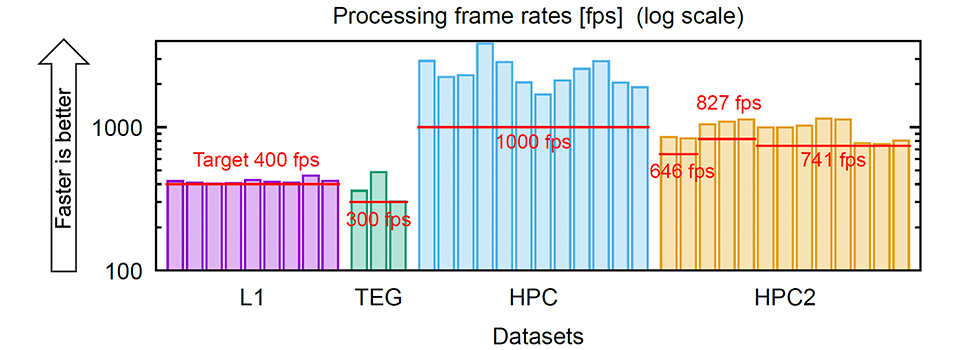
Putting all of these together, we processed 37 microscopy videos from multiple dataset groups listed in Table 1, and for all of them, we were able to achieve processing speeds that were higher than the respective video recording frame rates (Figure 3).
Compared with existing data processing methods for voltage imaging, the proposed method is more than 100 times faster than SGPMD-NMF[5] and 4-8 times faster than VolPy[6], and is the first to achieve real-time processing on a single desktop computer (whose specifications can be found in Table 2). The code is available on GitHub.
* Table can be scrolled horizontally.
|
Dataset group |
Animal |
Brain region |
Frame rate (fps) |
# videos |
Voltage indicator |
|---|---|---|---|---|---|
|
L1 |
Mouse |
L1 cortex |
400 |
9 |
Voltron[7] |
|
TEG |
Zebrafish |
Tegmentum |
300 |
3 |
Voltron[7] |
|
HPC |
Mouse |
Hippocampus |
1,000 |
12 |
paQuasAr3-s[8] |
|
HPC2 |
Mouse |
Hippocampus |
645 – 826 |
13 |
SomArchon[2] |
|
Component |
Specification |
|---|---|
|
CPU |
AMD Ryzen Threadripper 3960X (24 cores, 3.8 GHz) |
|
RAM |
DDR4 3200 MHz, 192 GB (6 channels x 32 GB) |
|
GPU |
2x NVIDIA GeForce RTX 2080 Ti |
|
SSD |
KIOXIA EXCERIA PRO 2 TB (NVMe™, PCIe® Gen 4 x4) |
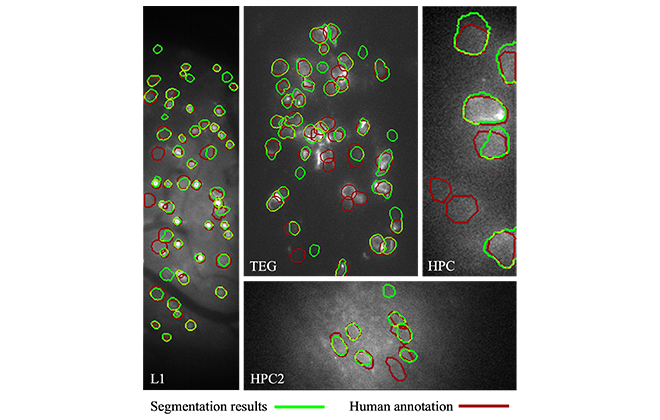
We evaluated the accuracy of the proposed neuron segmentation by comparing it with the ground truth accompanying the datasets. The agreement was about 80% on average. Some examples are shown in Figure 4. This accuracy is on par with or better than that of the existing methods. The ground truth human annotation was made by having multiple annotators manually draw neuron contours, and cross-check and revise them until everyone agreed. Because the images were noisy due to high-speed capture, the initial judgement by the individual annotators had only around 80% agreement with the consensus ground truth. Therefore, a segmentation accuracy of 80% can be viewed as being accurate enough to replace the manual labor of a single person.
Our pipeline leaves some data processing unsupported, including fixing neuron segmentation results that are different from what researchers expect, as well as further analysis of extracted voltage traces. Nevertheless, we believe it is useful for neuroscientists by being able to quickly present voltage traces of the majority of many captured neurons, and we hope it will help accelerate voltage imaging-based neuroscience. Kioxia will continue to contribute to cutting-edge research through high-speed data processing by leveraging SSD and memory technology.
This work was presented at the International Conference on Bioinformatics and Biomedicine (BIBM) 2023.
NVM Express, NVMe and NVMe-oF are registered or unregistered marks of NVM Express, Inc. in the United States and other countries.
PCI-SIG, PCI Express and PCIe are registered trademarks of PCI-SIG.
Other company names, product names and service names may be trademarks of third-party companies.
Reference
[1] Y. Bando et al. Real-time neuron segmentation for voltage imaging. IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 813–818, 2023.
[2] K. D. Piatkevich et al. Population imaging of neural activity in awake behaving mice. Nature 574, 413–417, 2019.
[3] J. P. Lewis. Fast template matching. Vision Interface, 120–123, 1995.
[4] O. Ronneberger, P. Fischer, and T. Brox. U-Net: Convolutional networks for biomedical image segmentation. Medical Image Computing and Computer-Assisted Intervention (MICCAI), 234–241, 2015.
[5] M. E. Xie et al. High-fidelity estimates of spikes and subthreshold waveforms from 1-photon voltage imaging in vivo. Cell Reports 35(1), 108954, 2021.
[6] C. Cai et al. VolPy: Automated and scalable analysis pipelines for voltage imaging datasets. PLoS Computational Biology 17(4), e1008806, 2021.
[7] A. S. Abdelfattah et al. Bright and photostable chemigenetic indicators for extended in vivo voltage imaging. Science 365(6454), 699–704, 2019.
[8] Y. Adam et al. Voltage imaging and optogenetics reveal behaviour-dependent changes in hippocampal dynamics. Nature 569, 413–417, 2019.

